Insights into the Discrepancy between Single Molecule Experiments *
|
Insights into the Discrepancy between Single Molecule Experiments * |
| Qian Zhou,Min Zhang,Yang-Tao Fan,Yu-Kang Wang,Lin Bao,Guang-Ju Zhao,Hu Chen,Yan-Hui Liu |
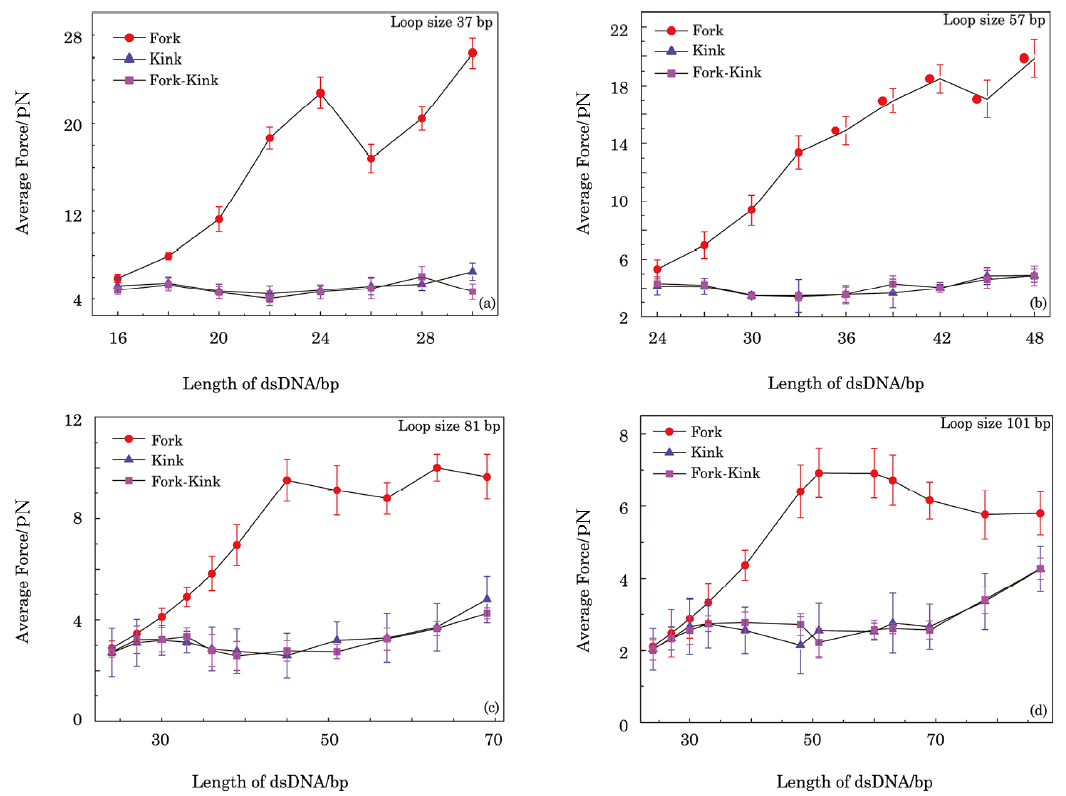
| Fig. 5 Forces extracted from simulations as function of contour length of dsDNA in sensor. Three conditions are considered in simulations, namely that without kink structure in dsDNA but with a fork at the junction of dsDNA and ssDNA, that with kink structure in dsDNA but without a fork at the junction of dsDNA and ssDNA, and that with kink structure in dsDNA and a fork at the junction of dsDNA and ssDNA. NN stacking interaction $\Delta\xi$ and the excited energy $\mu$ used in simulations are $2.7k_{B}T$ and $8k_{B}T$, respectively. |

|